lstm time series prediction in R
It turns out that deep learning, with all its power, can also be used for forecasting. Especially the LSTM (Long Short Term Memory) model, which proved to be useful while solving problems involving sequences with autocorrelation.
Long Short Term Memory networks are kind of Recurrent Neural Networks (RNN) that are capable of learning long-term dependencies. LSTM enables to persist long term states in addition to short term, which tradicional RNN’s have difficulty with. LSTMs are quite useful in time series prediction tasks involving autocorrelation, because of their ability to maintain state and recognize patterns over the length of the series.
Here I show how to implement forecasting LSTM model using R language.
Contents
HOW TO
LSTM model is available in the keras R package, which runs on top of the Tensorflow.
Before we start we need to install and load both of those:
|
1 2 3 4 |
library(keras) library(tensorflow) install_keras() install_tensorflow(version = "nightly") |
data preparation
For the purpose of this example I used the economics dataset, which can be found in the ggplot2 package. I want to predict the unemployment in the following 12 months (unemploy column).
|
1 2 |
library(ggplot2) head(economics) |
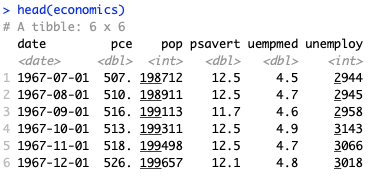
LSTM model requires to rescale the input data. Mean and standard deviation of the training dataset can be used as the scaling coefficients to scale both the training and testing data sets as well as the predicted values. This way we ensure that the scaling does not impact the model.
|
1 |
scale_factors <- c(mean(economics$unemploy), sd(economics$unemploy)) |
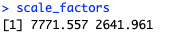
If you wish to train-test the model, you should start with data split. As I focused on creating the prediction, not accuracy of the model itself, I used full dataset for training.
|
1 2 3 |
scaled_train <- economics %>% dplyr::select(unemploy) %>% dplyr::mutate(unemploy = (unemploy - scale_factors[1]) / scale_factors[2]) |
LSTM algorithm creates predictions based on the lagged values. That means that it needs to look back to as many previous values as many points we wish to predict. As we want to do a 12 months forecast, we need to base each prediction on 12 data points.
For demonstration purpose let’s say our series has 10 data points [1, 2, 3, …, 10] and we want to predict 3 values. Then our training data looks like this:
|
1 2 3 4 5 |
[1, 2, 3] -> [4, 5, 6] [2, 3, 4] -> [5, 6, 7] [3, 4, 5] -> [6, 7, 8] [4, 5, 6] -> [7, 8, 9] [5, 6, 7] -> [8, 9, 10] |
Predictors and target take form of
|
1 2 3 |
3 4 5 6 7 6 7 8 9 10 X = [ 2 3 4 5 6 ] Y = [ 5 6 7 8 9 ] 1 2 3 4 5 4 5 6 7 8 |
Additionally keras LSTM expects specific tensor format of shape of a 3D array of the form [samples, timesteps, features] for predictors (X) and for target (Y) values:
- samples specifies the number of observations which will be processed in batches.
- timesteps tells us the number of time steps (lags). Or in other words how many units back in time we want our network to see.
- features specifies number of predictors (1 for univariate series and n for multivariate).
In case of predictors that translates to an array of dimensions: (nrow(data) – lag – prediction + 1, 12, 1), where lag = prediction = 12.
|
1 2 |
prediction <- 12 lag <- prediction |
We lag the data 11 times, so that each prediction is based on 12 values, and arrange lagged values into columns. Then we transform it into the desired 3D form.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 |
scaled_train <- as.matrix(scaled_train) # we lag the data 11 times and arrange that into columns x_train_data <- t(sapply( 1:(length(scaled_train) - lag - prediction + 1), function(x) scaled_train[x:(x + lag - 1), 1] )) # now we transform it into 3D form x_train_arr <- array( data = as.numeric(unlist(x_train_data)), dim = c( nrow(x_train_data), lag, 1 ) ) |
Basically every column is a lagged version of the previous one – the last one is lagged by 11 steps comparing to the first one.
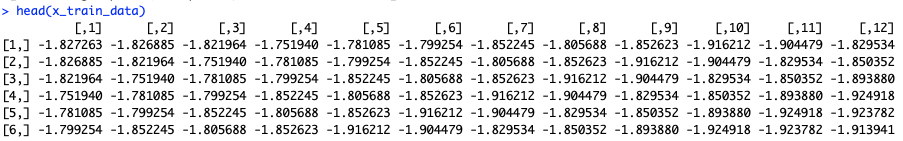
And final version:
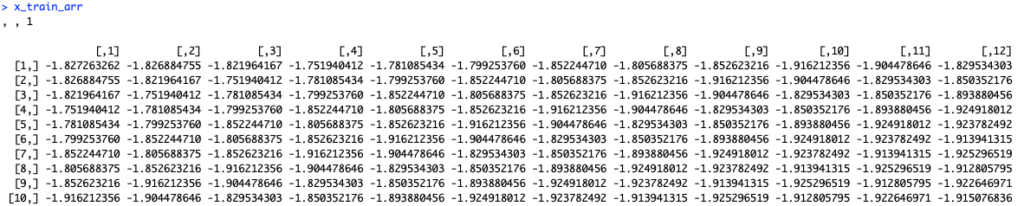
Data was turned into a 3D array. As we have only one predictor, last dimension equals to one.
Now we apply similar transformation for the Y values.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 |
y_train_data <- t(sapply( (1 + lag):(length(scaled_train) - prediction + 1), function(x) scaled_train[x:(x + prediction - 1)] )) y_train_arr <- array( data = as.numeric(unlist(y_train_data)), dim = c( nrow(y_train_data), prediction, 1 ) ) |
In the same manner we need to prepare input data for the prediction, which are in fact last 12 observations from our training set.
|
1 |
x_test <- economics$unemploy[(nrow(scaled_train) - prediction + 1):nrow(scaled_train)] |

We need to scale and transform it.
|
1 2 3 4 5 6 7 8 9 10 11 12 |
# scale the data with same scaling factors as for training x_test_scaled <- (x_test - scale_factors[1]) / scale_factors[2] # this time our array just has one sample, as we intend to perform one 12-months prediction x_pred_arr <- array( data = x_test_scaled, dim = c( 1, lag, 1 ) ) |

lstm prediction
We can build a LSTM model using the keras_model_sequential function and adding layers on top of that. The first LSTM layer takes the required input shape, which is the [samples, timesteps, features]. We set for both layers return_sequences = TRUE and stateful = TRUE. The second layer is the same with the exception of batch_input_shape, which only needs to be specified in the first layer.
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 |
lstm_model <- keras_model_sequential() lstm_model %>% layer_lstm(units = 50, # size of the layer batch_input_shape = c(1, 12, 1), # batch size, timesteps, features return_sequences = TRUE, stateful = TRUE) %>% # fraction of the units to drop for the linear transformation of the inputs layer_dropout(rate = 0.5) %>% layer_lstm(units = 50, return_sequences = TRUE, stateful = TRUE) %>% layer_dropout(rate = 0.5) %>% time_distributed(keras::layer_dense(units = 1)) |
You can also decide to try out different activation functions with activation parameter (hyperbolic tangent tanh is the default one).
Also choose loss function for the optimization, type of optimizer and metric for assessing the model performance. Info about different optimizers can he found here.
|
1 2 3 4 |
lstm_model %>% compile(loss = 'mae', optimizer = 'adam', metrics = 'accuracy') summary(lstm_model) |
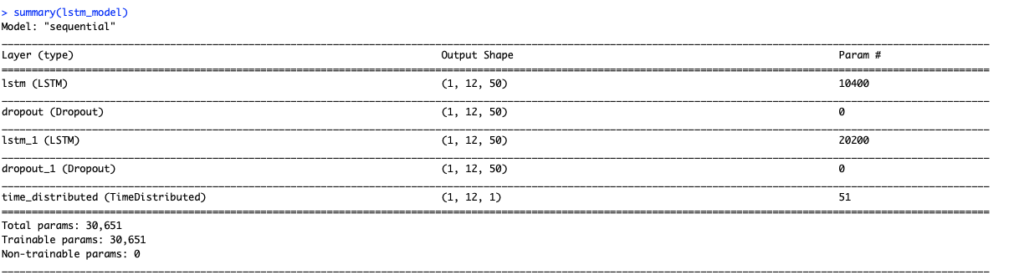

Next, we can fit our stateful LSTM. We set shuffle = FALSE to preserve sequences of time series.
|
1 2 3 4 5 6 7 8 |
lstm_model %>% fit( x = x_train_arr, y = y_train_arr, batch_size = 1, epochs = 20, verbose = 0, shuffle = FALSE ) |
And perform the prediction:
|
1 2 3 4 5 6 |
lstm_forecast <- lstm_model %>% predict(x_pred_arr, batch_size = 1) %>% .[, , 1] # we need to rescale the data to restore the original values lstm_forecast <- lstm_forecast * scale_factors[2] + scale_factors[1] |

LSTM model is more tricky than regular time series models, because you do not pass the explicit number of prediction points for the forecast. Instead you need to design your model in a way that it forecasts the desired number of periods. If you wish to predict more, you need to provide additional columns in your prediction set, containing values predicted for the previous periods. Following this example, predicting 13th month ahead requires “knowing” result for the 1st month ahead. The other option is to rebuild the model to predict 13 values instead of 12.
forecast object
As we have the values predicted, we can turn the results into the forecast object, as we would get if using the forecast package. That will allow i.e. to use the forecast::autoplot function to plot the results of the prediction. In order to do so, we need to define several objects that build a forecast object.
prediction on a train set
|
1 2 |
fitted <- predict(lstm_model, x_train_arr, batch_size = 1) %>% .[, , 1] |
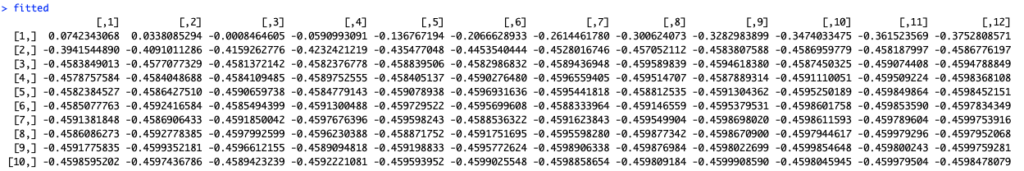
Prediction on a training set will provide us with 12 results for each input period. So we need to transform the data to get only one prediction per each date.
|
1 2 3 4 5 6 7 8 9 |
if (dim(fitted)[2] > 1) { fit <- c(fitted[, 1], fitted[dim(fitted)[1], 2:dim(fitted)[2]]) } else { fit <- fitted[, 1] } # additionally we need to rescale the data fitted <- fit * scale_factors[2] + scale_factors[1] nrow(fitted) # 562 |
Due to the fact that our forecast starts with 12 months offset, we need to provide artificial (or real) values for those months:
|
1 2 |
# I specify first forecast values as not available fitted <- c(rep(NA, lag), fitted) |
prediction in a form of ts object
We need to change the predicted values into a time series object.
|
1 2 3 4 5 |
lstm_forecast <- timetk::tk_ts(lstm_forecast, start = c(2015, 5), end = c(2016, 4), frequency = 12 ) |

input series
Additionally we need to transform the economics data into a time series object.
|
1 2 3 4 |
input_ts <- timetk::tk_ts(economics$unemploy, start = c(1967, 7), end = c(2015, 4), frequency = 12) |
forecast object
Finally we can define the forecast object:
|
1 2 3 4 5 6 7 8 9 10 |
forecast_list <- list( model = NULL, method = "LSTM", mean = lstm_forecast, x = input_ts, fitted = fitted, residuals = as.numeric(input_ts) - as.numeric(fitted) ) class(forecast_list) <- "forecast" |
Now we can easily plot the data:
|
1 |
forecast::autoplot(forecast_list) |
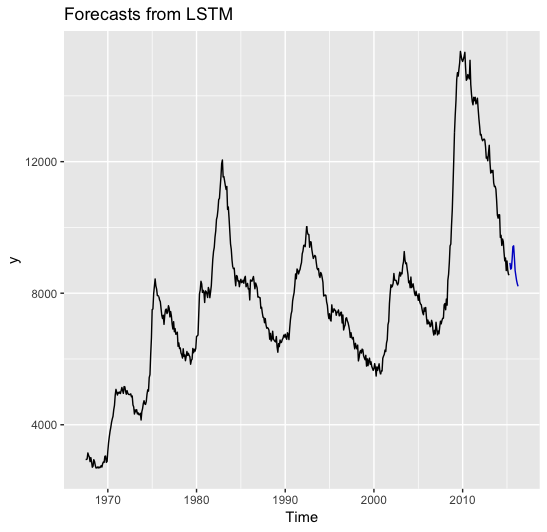
lstm prediction with regressors
Handling regressors in LSTM goes down to treating the series as multivariate instead of univariate. Let’s create some random regressors for this example:
|
1 |
reg <- 100 * runif(nrow(economics)) |

As with training set, we need to scale the regressors as well
|
1 2 3 4 5 6 7 8 9 10 |
scale_factors_reg <- list( mean = mean(reg), sd = sd(reg) ) scaled_reg <- (reg - scale_factors_reg$mean)/scale_factors_reg$sd # additionally 12 values for the forecast scaled_reg_prediction <- (reg[(length(reg) -12): length(reg)] - scale_factors_reg$mean) /scale_factors_reg$sd |
Now we can add them to the training data and transform to tensors as previously.
|
1 2 3 4 5 6 7 8 9 10 11 |
# combine training data with regressors x_train <- cbind(scaled_train, scaled_reg) x_train_data <- list() # transform the data into lagged columns for (i in 1:ncol(x_train)) { x_train_data[[i]] <- t(sapply( 1:(length(x_train[, i]) - lag - prediction + 1), function(x) x_train[x:(x + lag - 1), i] )) } |
This time we end up with 2 records in our list – input data in the exact same for as previously and regressors following the same logic.


Again we transform this list into a 3D array.
|
1 2 3 4 5 6 7 8 |
x_train_arr <- array( data = as.numeric(unlist(x_train_data)), dim = c( nrow(x_train_data[[1]]), lag, 2 ) ) |
We also need to modify the prediction data to include regressors in the same manner as for training.
|
1 2 3 4 5 6 7 8 9 10 11 12 |
# combine the data with regressors x_test_data <- c(x_test_scaled, scaled_reg_prediction) # transform to tensor x_pred_arr <- array( data = x_test_data, dim = c( 1, lag, 2 ) ) |

Rest of the modeling stays the same.
Many thanks for this article.
I am comfortable with R. Python’s system management and version compatibilities unsurmountable for many R users.
It is a lot easier to install TF and keras as root user as installing and configuring for non-admin user. As root user, everything ran on the first go.
I have got a project on sparse time series project and I am/was determined to stick to R whatever it takes, and your article was perfect in every manner.
Just few things I have to guess to instal forecast and timetk.
Thanks once again and really obliged.
Good evening!
In the section “lstm prediction with regressors”, where it says # combine training data with regressors
x_train <- cbind(scaled_train, scaled_reg)
x_train_data <- list()
Which ones are the vectors "scaled_train " and "scaled_reg"?
Thanks a lot for your help.
Hi,
“scaled_train” object you have in “data preparation” section, but generally it is a standardized time series.
“scaled_reg” is described in “lstm prediction with regressors”. Here again we have a standardize regressor for the prediction.
I’ve donde a lot of search to find tutorials for apply lstm in ts forecast, your is the best one.
Very helpful. I’ll be following your work.
Many thanks for this.
I’m trying to reproduce your work here, but am running into the following:
> lstm_model <- keras_model_sequential()
*** caught illegal operation ***
address 0x187ecf874, cause 'illegal trap'
Traceback:
1: py_module_import(module, convert = convert)
2: import(module)
3: doTryCatch(return(expr), name, parentenv, handler)
4: tryCatchOne(expr, names, parentenv, handlers[[1L]])
5: tryCatchList(expr, classes, parentenv, handlers)
6: tryCatch(import(module), error = clear_error_handler())
7: py_resolve_module_proxy(x)
8:
$.python.builtin.module(keras, "models")9: keras$models
10: keras_model_sequential()
Possible actions:
1: abort (with core dump, if enabled)
2: normal R exit
3: exit R without saving workspace
4: exit R saving workspace
Selection:
Any thoughts how to fix this? I'm not that familiar with Python unfortunately. Working on a MacBook Pro M1.
Thank you for sharing way too nice model!
I have a unsolved problem to develop a LSTM time series model.
I want to develop ‘multivariate LSTM prediction model’, but I don’t know how to coordinate the feature parameter to develop it…
And I don’t know how to prepare for multivariate datasets to apply into the model…
Can I get to know how to solve this problem?
This is super helpful, but any suggestions on how to generate predictions on new data into the future? I am guessing you’d have to predict one point at a time into the future, and use the preceding prediction as the lagged value?
Actually the example generates prediction of 12 months ahead. While building the model you need to specify the data lag equal the desired prediction period. Then as above each 12 observations predict next 12 observations, so last 12 points from your dataset is use to generate next 12. If you with to predict 24 periods then you can lag data 24 times or perform the 12 points prediction 12 times 🙂
Is it possible to do continues predictions using same model? Example, we predict 12 months in a year, use input from all of 2020 to predict 12 months of 2021. Then we use predicted values for 2021 as input to predict 12 months of 2022 and so on. Is that possible?
It is possible but usually not a good idea. Using predicted values to generate next predictions you will accumulate the prediction error. Each forecast comes with a confidence interval, so it is always better to use actual values to have more accurate prediction.
Hey, how one could try to build a model based on training and testing? is in your case, it’s full data. I tried but failed to run the codes. Like the codes, you used for 12 days ahead forecast. I want to divide the data into training and testing parts and then predict by using test data to check the model performance in terms of performance metrics, ME, MAE, MAPE, SAMPE etc.
Can the predictor data and target data have different number of columns or is it required that they both have the same number of columns?
Thank you for example. The first part with one regressor works but the second with two doesn’t. Would you please send me the whole code of the example or publish it. That is what I wrote:
lstm_model_m %
layer_lstm(units = 50, # size of the layer
batch_input_shape = c(1, 12, 2), # batch size, timesteps, features
return_sequences = TRUE,
stateful = TRUE) %>%
# fraction of the units to drop for the linear transformation of the inputs
layer_dropout(rate = 0.5) %>%
layer_lstm(units = 50,
return_sequences = TRUE,
stateful = TRUE) %>%
layer_dropout(rate = 0.5) %>%
time_distributed(keras::layer_dense(units = 1))
lstm_model %>%
compile(loss = ‘mae’, optimizer = ‘adam’, metrics = ‘accuracy’)
summary(lstm_model)
lstm_model %>% fit(
x = x_train_arr_m,
y = y_train_arr,
batch_size = 1,
epochs = 20,
verbose = 0,
shuffle = FALSE
)
I included features=2 in
batch_input_shape = c(1, 12, 2)
but response was
WARNING:tensorflow:Model was constructed with shape (1, 12, 1) for input KerasTensor(type_spec=TensorSpec(shape=(1, 12, 1), dtype=tf.float32, name=’lstm_7_input’), name=’lstm_7_input’, description=”created by layer ‘lstm_7_input'”), but it was called on an input with incompatible shape (None, 12, 2).
Thank you
Sure, hope that helps.
Hi I am facing this issue : please help
Error in loadNamespace(name): there is no package called ‘timetk’
Traceback:
1. timetk::tk_ts
2. getExportedValue(pkg, name)
3. asNamespace(ns)
4. getNamespace(ns)
5. loadNamespace(name)
6. withRestarts(stop(cond), retry_loadNamespace = function() NULL)
7. withOneRestart(expr, restarts[[1L]])
8. doWithOneRestart(return(expr), restart)
You need to have timetk package installed:
Thanks for posting this blog. It’s really helpful.
I have a question regarding your code.
y_train_data <- t(sapply(
(1 + lag):(length(scaled_train) – prediction + 1),
function(x) scaled_train[x:(x + prediction – 1)]
))
I think you don't need t() for this matrix?
Have a great day 🙂
I need the data to match training set – so having 12 columns.
I’m having problems with the plot of the results, while applying:
forecast::autoplot(forecast_list)
is returning this warning error message:
> autoplot(forecast_list)
Error:
datamust be a data frame, or other object coercible byfortify(), not a numeric vector.Run
rlang::last_error()to see where the error occurred.Anyone has face the same problem?
very helpful and informative.
Its really helpful posting, Thank you.
in your code, lag period is equal to prediction period,
but when i set lag != prediction period,
lstm calls error. like this
Error in py_call_impl(callable, dots$args, dots$keywords) :
ValueError: in user code:
i want to predict 5 time sequence based on last 90 time sequence.
so that would be setting lag =90 , prediction =5
pls help me :))
thank you
Generally in LSTMs you do not have such a limitation, but for the time series prediction it is required. You need to build a network capable of predicting given number of points and for that it shall use rolling window of the same length.
So for 5 point prediction you should use lag=5, but build on the whole 90 points sequence.
please could you explain this section again
we lag the data 11 times and arrange that into columns
x_train_data <- t(sapply(
1:(length(scaled_train) – lag – prediction + 1),
function(x) scaled_train[x:(x + lag – 1), 1]
))
i went through your work and i am ok with everything but the above section on univariate
time series is still unclear to me.
i have a univariate time time series of period 12 and i want to be able to predict the next 6months , 8 months and 12 months data.
You can either build different models for different time spans, or use the one with 12 predictions to obtain just 6 or 8 points (ignoring the rest).
Hi i would to know how to forecast more than 12 months ( 20, 26, 14,…) with the same model for univariate data.
Thanks
Thanks for this blog.
I would need help with the process with the daily data.
Should work in the same way. Use daily data and properly lag them to get the desired prediction.
Hi Alice,
I wrote bunch of functions which somehow automates your work. It works fine with weekly, monthly, quarterly and anually univariate time-series.
Please check out:
https://rpubs.com/pawel-wieczynski/891765
Thanks for sharing!
Can you please share the R-codes for monthly, quarterly and annually univariate time-series forecasting.
i,
ow can we find accuracy?
I am unable to find prediction on daily basis.
Thanks for this method, I have a question I got this error
Error in normalizePath(path.expand(path), winslash, mustWork) :
path[1]=”C:/PROGRA~3/MINICO~1/Scripts/conda.exe”: The system cannot find the path specified
Please help me
* Installing Miniconda — please wait a moment …
Error: miniconda installation failed [exit code 2]
I get this error while installing Keras by this command > install_keras()
Any idea how I can proceed ?
Hi,I would like to know,I have Minute wise Time series data.I want predict for upcoming onedaay(1440) blocks,so i fixed my lag as 1440 and i built the model.Train and test model accuracy iis pretty much accurate.But when i compare to forecasted day’s actual and predicted values its not accurate,Which like i spli the data first 19 days as train and i passes 20th data as test and predicted for 21.When i compared 20th actual and forecasted data its accurate.But when i compared with 21(the actual forecasyed day) its coming less.So what would be the cause????
Help me……………!
I am a bit confused about how I would calculate error metrics in this prediction.
I guess I should divide the data into training, testing and validation datasets. In this way, I would train the model in the training, make predictions on the testing and check these predictions against the validation dataset.
Is it like that?
Yes 🙂
Muhammad Naeem, generally I would suggest to have the testing data of the same length as the planned prediction. So if you plan to predict 3 points ahead, use 3 points as test set. Then you can use rolling window of 1 period to backtest even further (using always ordered observations (1, t) as training and (t+1, t+3) as testing) That would give you more accurate overview.
timetk never want to be installed in r studio.
why you use complicated libraries please.
it keeps asking to update other packages but at the end never want to get installed
Thank you for such a useful article!
Nevertheless, I have 51 dataset to which I would like to apply lstm and make the predictions, and I don’t quite understand how to do it at the same time without the need to run lstm for 51 time.
I tried to do it via the loop for, so to apply lstm to each element of the list of 51 elements, but it seems lstm_model object can not be a list.
More in detail, I tried starting from
lstm_model=list()
for(i in 1:n){
lstm_model[[i]] <- keras_model_sequential()
…. and so on adding the index i to each of the lstm_model and trying to run lstm in a loop going through the list of the 51 datasets.
}
but it does not work.
How would you suggest to solve this problem?
Thank you a lot in advance!
Lyubov
Sorry, actually I have just solved this issue. Thank you for such a useful content!
Lyubov
But I am still not sure why for the purpose of predictions the predict function does not accept the fitted object. It works only with the compiled object, otherwise I get the error
Error in UseMethod(“predict”) :
no applicable method for ‘predict’ applied to an object of class “keras_training_history”
Shouldn’t the predict function have the fitted model as an object in order to make the predictions?
Thank you
Hi! I am not quite understanding how you are making predictions if before you run the model you already have the test period rescaled. How should the model be set in the case I don’t have the test period at all?
x_test <- economics$unemploy[(nrow(scaled_train) – prediction + 1):nrow(scaled_train)]
Here you are considering the scaled_train which represents the full period. But in my case scaled_train is only the training period, is x_test defined in the same way?
Thank you
As described at the top in this model, you use previous observations to predict next. So in order to have the prediction you need to supply prediction function with a few last observations, which I called “x_test”.
how to visualisation withthis method?
hi, I keep encountering this error:
> lstm_model <- keras_model_sequential()
Error in python_config_impl(python) :
Error 1 occurred running /usr/bin/python3:
can you help? Thanks!